Intro to Threads and Processes in Python

Competing in Kaggle’s Understanding the Amazon from Space competition, I started timing various parts of my code to see if I could speed things up. Speed is critical in Kaggle. Ranking well often requires trying hundreds of architectural and hyper-parameter combinations. Shaving 10 seconds off an epoch that lasts 1 minute is a huge win.
To my surprise, I found data augmentation was my biggest bottleneck. The methods I used — rotations, flips, zooms and crops — relied on Numpy and ran on the CPU. Numpy uses parallel processing in some cases and Pytorch’s data loaders do as well, but I was running 3–5 experiments at a time and each experiment was doing its own augmentation. This seemed inefficient and I was curious to see if I could speed things up with parallel processing.
What is parallel processing?
Basically doing two things at the same time, either running code simultaneously on different CPUs, or running code on the same CPU and achieving speedups by taking advantage of “wasted” CPU cycles while your program is waiting for external resources — file loading, API calls.
As an example, here is a “normal” program. It downloads a list of URLs one at a time using a single thread.
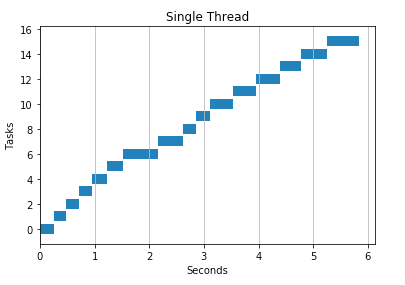
Here is the same program using 2 threads. It divides urls among threads giving us nearly a 2x speedup.
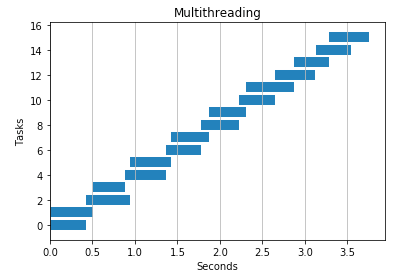
If you’re curious how to generate these charts and what they mean you can find the code here, but to briefly summarize:
- Add a timer inside your function and return its start and stop time
URLS = [url1, url2, url3, ...]
def download(url, base):
start = time.time() - base
resp = urlopen(url)
stop = time.time() - base
return start,stop
2. To visualize a single thread, run your function multiple times and store the start and stop times
results = [download(url, 1) for url in URLS]
3. Transpose the resulting array of [start, stop] times and plot a bar chart
def visualize_runtimes(results):
start,stop = np.array(results).T
plt.barh(range(len(start)), stop-start, left=start)
plt.grid(axis=’x’)
plt.ylabel("Tasks")
plt.xlabel("Seconds")
Charts for multiple threads can be generated the same way. The methods in Python’s concurrency library return an array of results.
Process vs Thread
A process is an instance of program (e.g. Jupyter notebook, Python interpreter). Processes spawn threads (sub-processes) to handle subtasks like reading keystrokes, loading HTML pages, saving files. Threads live inside processes and share the same memory space.
Example: Microsoft Word
When you open Word, you create a process. When you start typing, the process spawns threads: one to read keystrokes, another to display text, one to autosave your file, and yet another to highlight spelling mistakes. By spawning multiple threads, Microsoft takes advantage of idle CPU time (waiting for keystrokes or files to load) and makes you more productive.
Process
- Created by the operating system to run programs
- Processes can have multiple threads
- Two processes can execute code simultaneously in the same python program
- Processes have more overhead than threads as opening and closing processes takes more time
- Sharing information between processes is slower than sharing between threads as processes do not share memory space. In python they share information by pickling data structures like arrays which requires IO time.
Thread
- Threads are like mini-processes that live inside a process
- They share memory space and efficiently read and write to the same variables
- Two threads cannot execute code simultaneously in the same python program (although there are workarounds*)
CPU vs Core
The CPU, or processor, manages the fundamental computational work of the computer. CPUs have one or more cores, allowing the CPU to execute code simultaneously.
With a single core, there is no speedup for CPU-intensive tasks (e.g. loops, arithmetic). The OS switches back and forth between tasks executing each one a little bit at a time. This is why for small operations (e.g. downloading a few images), multitasking can sometimes hurt your performance. There is overhead associated with launching and maintaining multiple tasks.
Python’s GIL problem
CPython (the standard python implementation) has something called the GIL (Global Interpreter Lock), which prevent two threads from executing simultaneously in the same program. Some people are upset by this, while others fiercely defend it. There are workarounds, however, and libraries like Numpy bypass this limitation by running external code in C.
When to use threads vs processes?
- Processes speed up Python operations that are CPU intensive because they benefit from multiple cores and avoid the GIL.
- Threads are best for IO tasks or tasks involving external systems because threads can combine their work more efficiently. Processes need to pickle their results to combine them which takes time.
- Threads provide no benefit in python for CPU intensive tasks because of the GIL.
*For certain operations like Dot Product, Numpy works around Python’s GIL and executes code in parallel.
Parallel processing examples
Python’s concurrent.futures library is surprisingly pleasant to work with. Simply pass in your function, a list of items to work on, and the number of workers. In the next few sections, I walk through various experiments I ran to learn more about when to use threads vs processing.
def multithreading(func, args,
workers):
with ThreadPoolExecutor(workers) as ex:
res = ex.map(func, args)
return list(res)
def multiprocessing(func, args,
workers):
with ProcessPoolExecutor(work) as ex:
res = ex.map(func, args)
return list(res)
API calls
I found threads work better for API calls and observed speedups over serial processing and multiprocessing.
def download(url):
try:
resp = urlopen(url)
except Exception as e:
print ('ERROR: %s' % e)
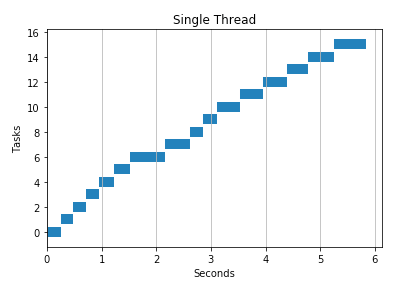
2 threads
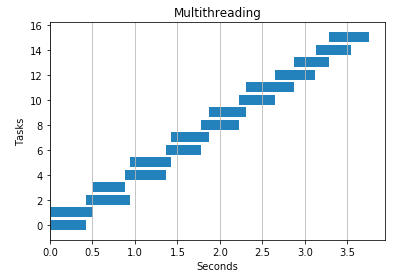
4 threads
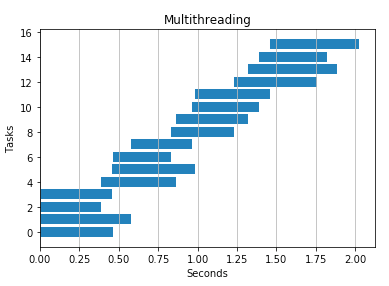
2 processes
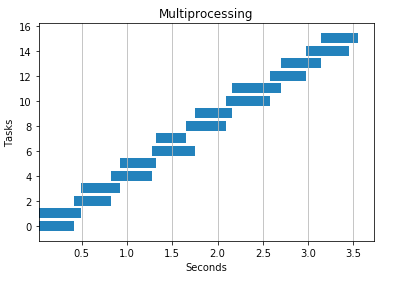
4 processes
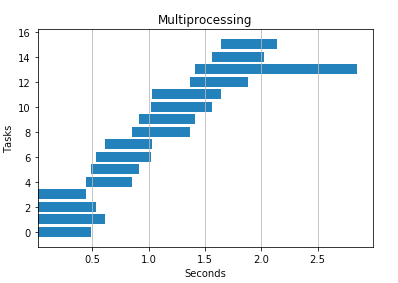
IO Heavy Task
I passed in a bunch of huge text strings to see how write performance differed. Threads seemed to win here, but multiprocessing also improved runtime.
def io_heavy(text):
f = open('output.txt', 'wt', encoding='utf-8')
f.write(text)
f.close()
Serial
%timeit -n 1 [io_heavy(TEXT,1) for i in range(N)]
>> 1 loop, best of 3: 1.37 s per loop
4 threads
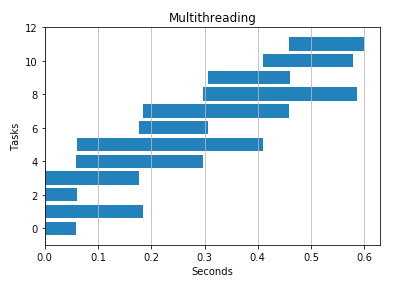
4 processes

CPU Intensive
Multiprocessing won the day here as expected. Processes avoid the GIL and execute code simultaneously on multiple cores.
def cpu_heavy(n):
count = 0
for i in range(n):
count += i
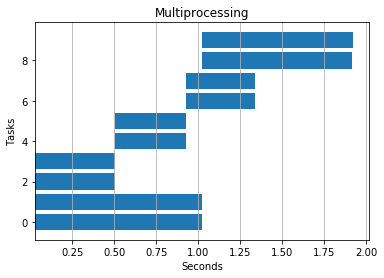
Serial: 4.2 seconds
4 threads: 6.5 seconds
4 processes: 1.9 seconds
Numpy Dot Product
As expected, I saw no benefit adding threads or processes to this code. Numpy executes external C code behind the scenes and thus evades the GIL.
def dot_product(i, base):
start = time.time() - base
res = np.dot(a,b)
stop = time.time() - base
return start,stop
Serial: 2.8 seconds
2 threads: 3.4 seconds
2 processes: 3.3 seconds
Here’s a notebook you can use to replicate these experiments on your own.
Resources
Here are the articles I enjoyed and referenced while I explored this topic. In particular I want to call out Nathan Grigg’s blog post, which gave me the idea for the visualizations.
☞ Python for Beginners: Become a Certified Python Developer
☞ Introduction to Python for Beginners
☞ The Python 3 Bible™ | Go from Beginner to Advanced in Python
☞ Complete Python Bootcamp: Go from zero to hero in Python
Suggest:
☞ Python Tutorials for Beginners - Learn Python Online
☞ What is Python and Why You Must Learn It in [2019]
☞ Complete Python Tutorial for Beginners (2019)
☞ Learn Python in 12 Hours | Python Tutorial For Beginners
☞ Python Tutorial for Beginners [Full Course] 2019
☞ Python Programming Tutorial | Full Python Course for Beginners 2019